Genomiverse
Uncovering Diverse Microbial Mechanisms via AI
Home DNAProteinBinding MicrobiomeConnectome JeffsKnowledgeGraph Publications News AnniBot Our Team Contact Us Game

What do we do?
- AI models to predict how proteins interact with DNA and ligands (coming soon).
- MicrobiomeConnectome: knowledge graph for candidate drug prediction!
How we do it?
We build AI from biological principles and generate training data from data we already have:
- >3M microbial genomes and >30K microbial species
- 143K microbiome literature
Who are we?
We are a team of microbiologists and software engineers previously from MIT and Google.
Other tools
- DNA sequence aligner: Mapper
- Fast variant detector: QuickVariants
- AMR risk rank detector: arg_ranker
- Our github

Highlighted discoveries & publications
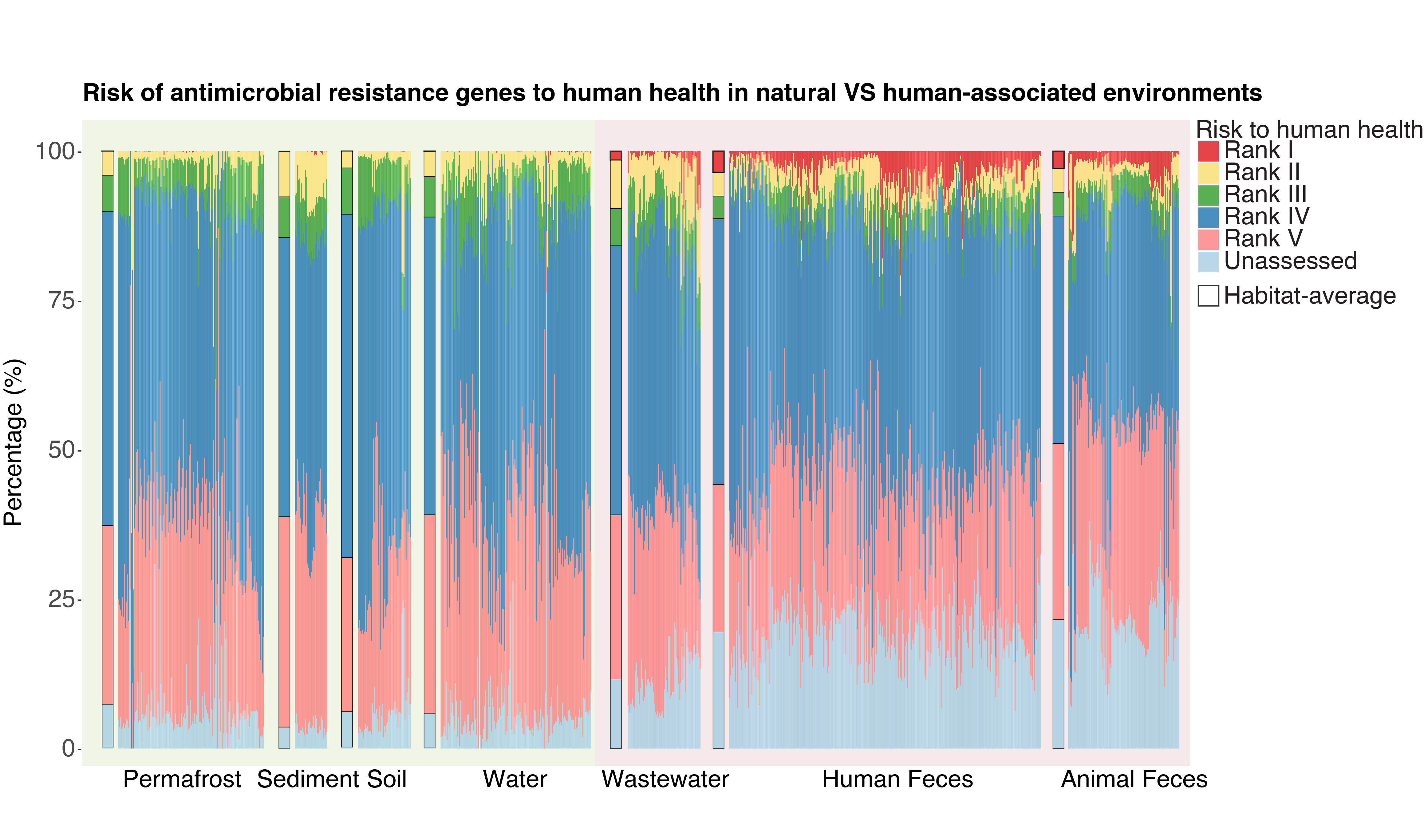
Antibiotic Resistance Gene (ARG) Risk
Antibiotic resistance genes (ARGs) are everywhere, but enrichment in human-associated environments is considered a requirement to be the highest risk (Rank I - red bars).
Read Publication →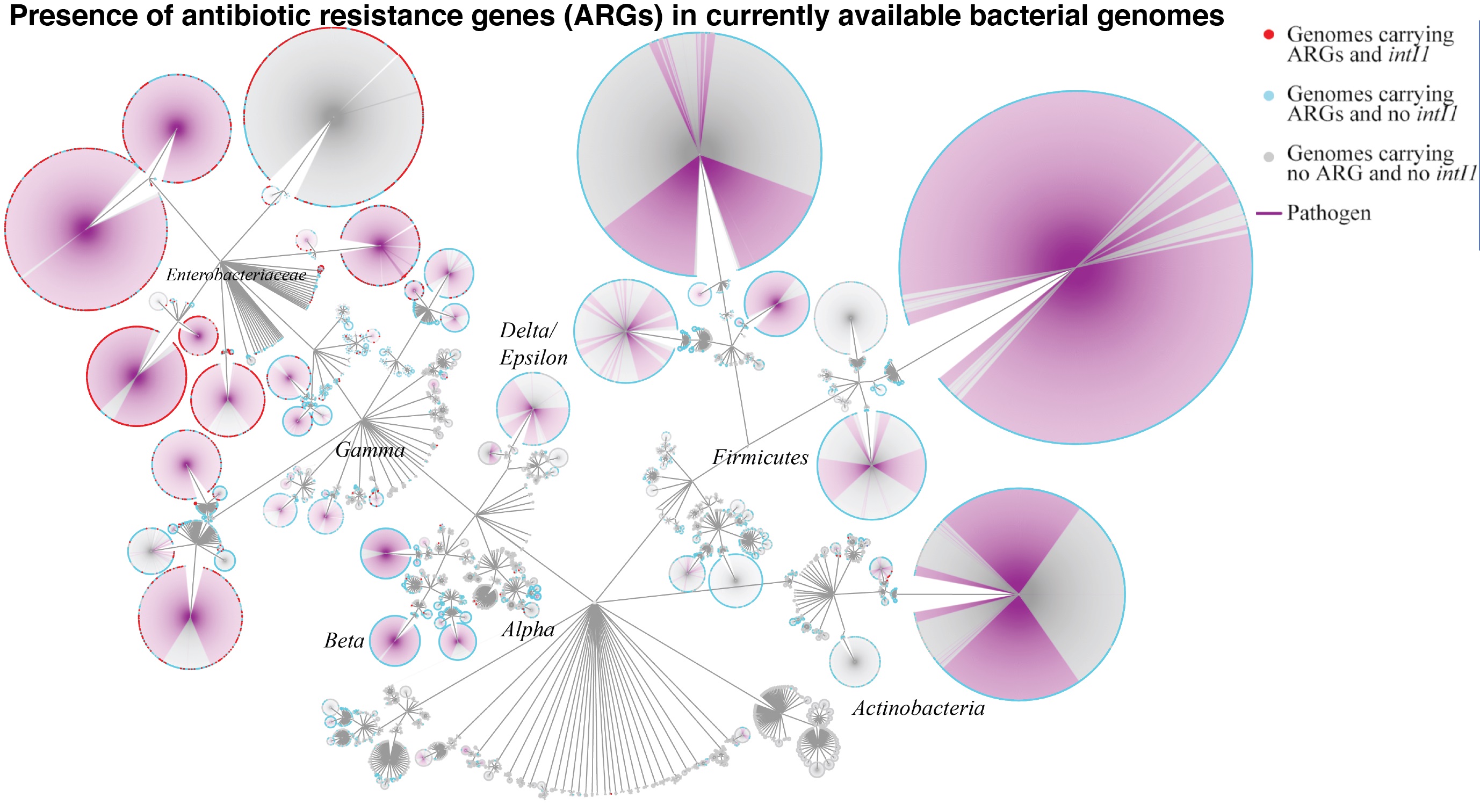
ARG Prevalence & Spread
ARGs are highly prevalent in bacterial genomes. Specifically, ARGs in Gammaproteobacteria are spreading by horizontal gene transfer via class 1 integrons (i.e. intI1).
Read Publication →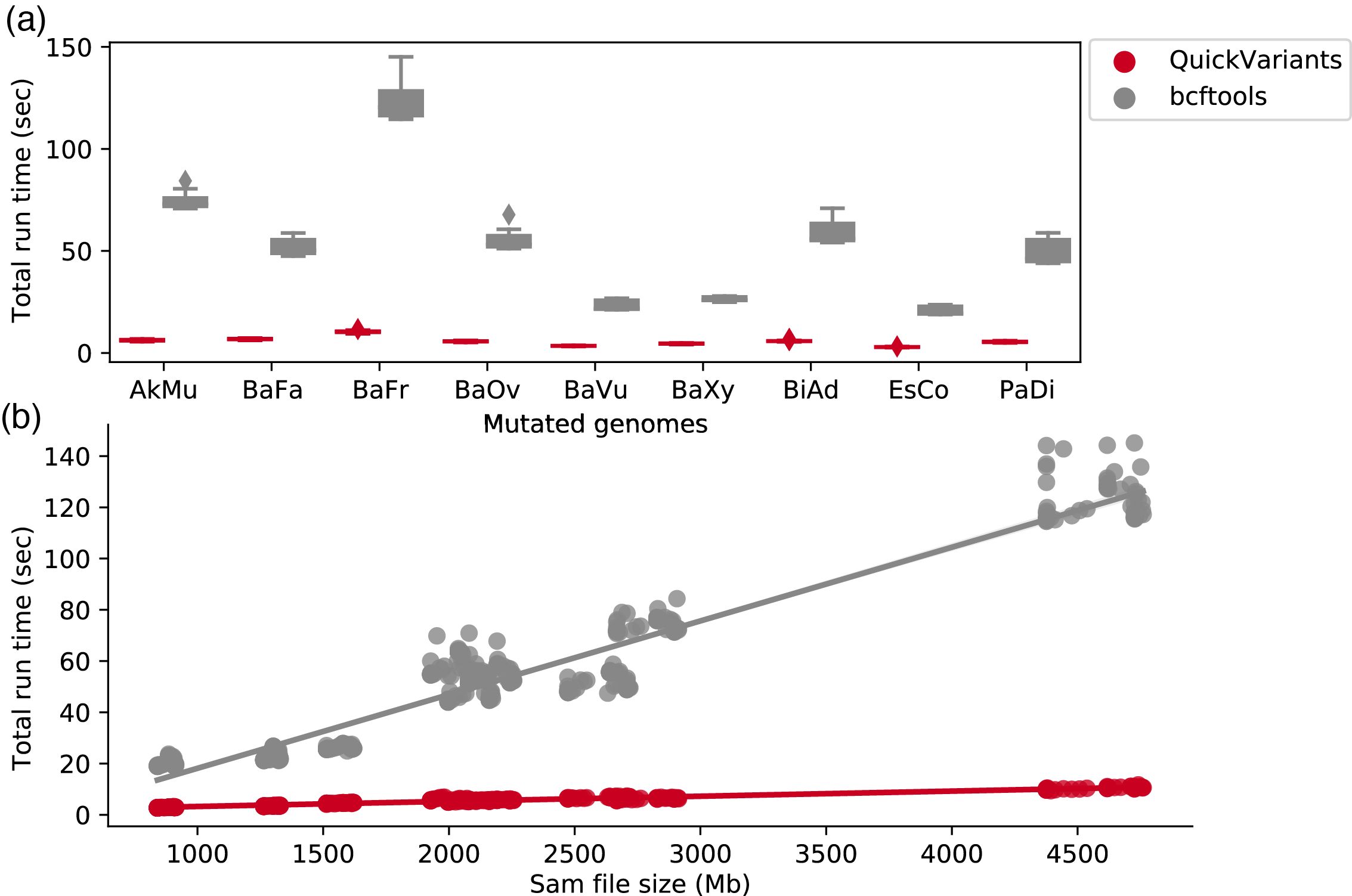
QuickVariants - Variant Detection
A novel algorithm (QuickVariants) for genetic variant identification that is 9x faster than existing tools, while producing fewer false positives and false negatives.
Read Publication →
Mapper - Sequence Aligner
Our tool, Mapper, dynamically incorporates k-mers of various sizes and gaps (gapped x-mers) to significantly improve sequence alignment quality.
Read Publication →
Gut CRISPR Immunology
Gut microbiomes share CRISPR systems through horizontal gene transfer to dynamically update their phage defense. Selected as Cover Image and reported by MIT News.
Read Publication →
CRISPR Adaptation
A highly prevalent CRISPR array in Bifidobacterium longum spreading via horizontal gene transfer (HGT), with six spacers found in 15 persons from the United States and Europe
Read Publication →